Dataset Details
C1 CAGE and RNA-seq quantification of single-cell gene expression during TGF-β stimulation of A549 cells.
C1 CAGE is a method to quantify single-cell gene expression by transcriptome sequencing on the Illumina platform. This method generates random-primed cDNAs, which allows detection of 5'-end transcripts to map transcription start sites (TSS) at single-nucleotide resolution. This method is performed in the microfluidics chips of the Fluidigm C1 system. The single cell expression data is derived from human A549 adenocarcinomic human alveolar basal epithelial cells stimulated with TGF-β for 0, 6 and 24 h. We used the "Rev A" and "Rev B" version of C1 CAGE, where ERCC spikes are diluted 200 and 20,000 times, respectively, and the standard C1 RNA-seq protocol (PN 100-7168 I1). The stimulations were ran in triplicates, which were multiplexed to control for chip bias using a color-coding strategy: cells stimulated at 0, 6 and 24 hrs were stained with different calcein dyes and pooled prior to loading into the C1 system. Raw images of capture sites were taken using Cellomics ArrayScan VTI High Content Analysis Reader (single images) or IN Cell Analyzer 6000 (z-stack).
Cellomics Images
Bulk Download| Accession Number | Brightfield Image | Green Fluorescence Image | Red Fluorescence Image |
|---|---|---|---|
| 1772-064-101_A01 |  |
 |
 |
| 1772-064-101_A02 |  |
 |
 |
| 1772-064-101_A03 |  |
 |
 |
| 1772-064-101_A04 |  |
 |
 |
| 1772-064-101_A05 |  |
 |
 |
| 1772-064-101_A06 |  |
 |
 |
| 1772-064-101_A07 |  |
 |
 |
| 1772-064-101_A08 |  |
 |
 |
| 1772-064-101_A09 |  |
 |
 |
| 1772-064-101_A10 |  |
 |
 |
| 1772-064-101_A11 |  |
 |
 |
| 1772-064-101_A12 |  |
 |
 |
| 1772-064-101_B01 |  |
 |
 |
| 1772-064-101_B02 |  |
 |
 |
| 1772-064-101_B03 |  |
 |
 |
| 1772-064-101_B04 |  |
 |
 |
| 1772-064-101_B05 |  |
 |
 |
| 1772-064-101_B06 |  |
 |
 |
| 1772-064-101_B07 |  |
 |
 |
| 1772-064-101_B08 |  |
 |
 |
| 1772-064-101_B09 |  |
 |
 |
| 1772-064-101_B10 |  |
 |
 |
| 1772-064-101_B11 |  |
 |
 |
| 1772-064-101_B12 |  |
 |
 |
| 1772-064-101_C01 |  |
 |
 |
| 1772-064-101_C02 |  |
 |
 |
| 1772-064-101_C03 |  |
 |
 |
| 1772-064-101_C04 |  |
 |
 |
| 1772-064-101_C05 |  |
 |
 |
| 1772-064-101_C06 |  |
 |
 |
| 1772-064-101_C07 |  |
 |
 |
| 1772-064-101_C08 |  |
 |
 |
| 1772-064-101_C09 |  |
 |
 |
| 1772-064-101_C10 |  |
 |
 |
| 1772-064-101_C11 |  |
 |
 |
| 1772-064-101_C12 |  |
 |
 |
| 1772-064-101_D01 |  |
 |
 |
| 1772-064-101_D02 |  |
 |
 |
| 1772-064-101_D03 |  |
 |
 |
| 1772-064-101_D04 |  |
 |
 |
| 1772-064-101_D05 |  |
 |
 |
| 1772-064-101_D06 |  |
 |
 |
| 1772-064-101_D07 |  |
 |
 |
| 1772-064-101_D08 |  |
 |
 |
| 1772-064-101_D09 |  |
 |
 |
| 1772-064-101_D10 |  |
 |
 |
| 1772-064-101_D11 |  |
 |
 |
| 1772-064-101_D12 |  |
 |
 |
| 1772-064-101_E01 |  |
 |
 |
| 1772-064-101_E02 |  |
 |
 |
| 1772-064-101_E03 |  |
 |
 |
| 1772-064-101_E04 |  |
 |
 |
| 1772-064-101_E05 |  |
 |
 |
| 1772-064-101_E06 |  |
 |
 |
| 1772-064-101_E07 |  |
 |
 |
| 1772-064-101_E08 |  |
 |
 |
| 1772-064-101_E09 |  |
 |
 |
| 1772-064-101_E10 |  |
 |
 |
| 1772-064-101_E11 |  |
 |
 |
| 1772-064-101_E12 |  |
 |
 |
| 1772-064-101_F01 |  |
 |
 |
| 1772-064-101_F02 |  |
 |
 |
| 1772-064-101_F03 |  |
 |
 |
| 1772-064-101_F04 |  |
 |
 |
| 1772-064-101_F05 |  |
 |
 |
| 1772-064-101_F06 |  |
 |
 |
| 1772-064-101_F07 |  |
 |
 |
| 1772-064-101_F08 |  |
 |
 |
| 1772-064-101_F09 |  |
 |
 |
| 1772-064-101_F10 |  |
 |
 |
| 1772-064-101_F11 |  |
 |
 |
| 1772-064-101_F12 |  |
 |
 |
| 1772-064-101_G01 |  |
 |
 |
| 1772-064-101_G02 |  |
 |
 |
| 1772-064-101_G03 |  |
 |
 |
| 1772-064-101_G04 |  |
 |
 |
| 1772-064-101_G05 |  |
 |
 |
| 1772-064-101_G06 |  |
 |
 |
| 1772-064-101_G07 |  |
 |
 |
| 1772-064-101_G08 |  |
 |
 |
| 1772-064-101_G09 |  |
 |
 |
| 1772-064-101_G10 |  |
 |
 |
| 1772-064-101_G11 |  |
 |
 |
| 1772-064-101_G12 |  |
 |
 |
| 1772-064-101_H01 |  |
 |
 |
| 1772-064-101_H02 |  |
 |
 |
| 1772-064-101_H03 |  |
 |
 |
| 1772-064-101_H04 |  |
 |
 |
| 1772-064-101_H05 |  |
 |
 |
| 1772-064-101_H06 |  |
 |
 |
| 1772-064-101_H07 |  |
 |
 |
| 1772-064-101_H08 |  |
 |
 |
| 1772-064-101_H09 |  |
 |
 |
| 1772-064-101_H10 |  |
 |
 |
| 1772-064-101_H11 |  |
 |
 |
| 1772-064-101_H12 |  |
 |
 |
| 1772-064-102_A01 |  |
 |
 |
| 1772-064-102_A02 |  |
 |
 |
| 1772-064-102_A03 |  |
 |
 |
| 1772-064-102_A04 |  |
 |
 |
| 1772-064-102_A05 |  |
 |
 |
| 1772-064-102_A06 |  |
 |
 |
| 1772-064-102_A07 |  |
 |
 |
| 1772-064-102_A08 |  |
 |
 |
| 1772-064-102_A09 |  |
 |
 |
| 1772-064-102_A10 |  |
 |
 |
| 1772-064-102_A11 |  |
 |
 |
| 1772-064-102_A12 |  |
 |
 |
| 1772-064-102_B01 |  |
 |
 |
| 1772-064-102_B02 |  |
 |
 |
| 1772-064-102_B03 |  |
 |
 |
| 1772-064-102_B04 |  |
 |
 |
| 1772-064-102_B05 |  |
 |
 |
| 1772-064-102_B06 |  |
 |
 |
| 1772-064-102_B07 |  |
 |
 |
| 1772-064-102_B08 |  |
 |
 |
| 1772-064-102_B09 |  |
 |
 |
| 1772-064-102_B10 |  |
 |
 |
| 1772-064-102_B11 |  |
 |
 |
| 1772-064-102_B12 |  |
 |
 |
| 1772-064-102_C01 |  |
 |
 |
| 1772-064-102_C02 |  |
 |
 |
| 1772-064-102_C03 |  |
 |
 |
| 1772-064-102_C04 |  |
 |
 |
| 1772-064-102_C05 |  |
 |
 |
| 1772-064-102_C06 |  |
 |
 |
| 1772-064-102_C07 |  |
 |
 |
| 1772-064-102_C08 |  |
 |
 |
| 1772-064-102_C09 |  |
 |
 |
| 1772-064-102_C10 |  |
 |
 |
| 1772-064-102_C11 |  |
 |
 |
| 1772-064-102_C12 |  |
 |
 |
| 1772-064-102_D01 |  |
 |
 |
| 1772-064-102_D02 |  |
 |
 |
| 1772-064-102_D03 |  |
 |
 |
| 1772-064-102_D04 |  |
 |
 |
| 1772-064-102_D05 |  |
 |
 |
| 1772-064-102_D06 |  |
 |
 |
| 1772-064-102_D07 |  |
 |
 |
| 1772-064-102_D08 |  |
 |
 |
| 1772-064-102_D09 |  |
 |
 |
| 1772-064-102_D10 |  |
 |
 |
| 1772-064-102_D11 |  |
 |
 |
| 1772-064-102_D12 |  |
 |
 |
| 1772-064-102_E01 |  |
 |
 |
| 1772-064-102_E02 |  |
 |
 |
| 1772-064-102_E03 |  |
 |
 |
| 1772-064-102_E04 |  |
 |
 |
| 1772-064-102_E05 |  |
 |
 |
| 1772-064-102_E06 |  |
 |
 |
| 1772-064-102_E07 |  |
 |
 |
| 1772-064-102_E08 |  |
 |
 |
| 1772-064-102_E09 |  |
 |
 |
| 1772-064-102_E10 |  |
 |
 |
| 1772-064-102_E11 |  |
 |
 |
| 1772-064-102_E12 |  |
 |
 |
| 1772-064-102_F01 |  |
 |
 |
| 1772-064-102_F02 |  |
 |
 |
| 1772-064-102_F03 |  |
 |
 |
| 1772-064-102_F04 |  |
 |
 |
| 1772-064-102_F05 |  |
 |
 |
| 1772-064-102_F06 |  |
 |
 |
| 1772-064-102_F07 |  |
 |
 |
| 1772-064-102_F08 |  |
 |
 |
| 1772-064-102_F09 |  |
 |
 |
| 1772-064-102_F10 |  |
 |
 |
| 1772-064-102_F11 |  |
 |
 |
| 1772-064-102_F12 |  |
 |
 |
| 1772-064-102_G01 |  |
 |
 |
| 1772-064-102_G02 |  |
 |
 |
| 1772-064-102_G03 |  |
 |
 |
| 1772-064-102_G04 |  |
 |
 |
| 1772-064-102_G05 |  |
 |
 |
| 1772-064-102_G06 |  |
 |
 |
| 1772-064-102_G07 |  |
 |
 |
| 1772-064-102_G08 |  |
 |
 |
| 1772-064-102_G09 |  |
 |
 |
| 1772-064-102_G10 |  |
 |
 |
| 1772-064-102_G11 |  |
 |
 |
| 1772-064-102_G12 |  |
 |
 |
| 1772-064-102_H01 |  |
 |
 |
| 1772-064-102_H02 |  |
 |
 |
| 1772-064-102_H03 |  |
 |
 |
| 1772-064-102_H04 |  |
 |
 |
| 1772-064-102_H05 |  |
 |
 |
| 1772-064-102_H06 |  |
 |
 |
| 1772-064-102_H07 |  |
 |
 |
| 1772-064-102_H08 |  |
 |
 |
| 1772-064-102_H09 |  |
 |
 |
| 1772-064-102_H10 |  |
 |
 |
| 1772-064-102_H11 |  |
 |
 |
| 1772-064-102_H12 |  |
 |
 |
| 1772-064-105_A01 |  |
 |
 |
| 1772-064-105_A02 |  |
 |
 |
| 1772-064-105_A03 |  |
 |
 |
| 1772-064-105_A04 |  |
 |
 |
| 1772-064-105_A05 |  |
 |
 |
| 1772-064-105_A06 |  |
 |
 |
| 1772-064-105_A07 |  |
 |
 |
| 1772-064-105_A08 |  |
 |
 |
| 1772-064-105_A09 |  |
 |
 |
| 1772-064-105_A10 |  |
 |
 |
| 1772-064-105_A11 |  |
 |
 |
| 1772-064-105_A12 |  |
 |
 |
| 1772-064-105_B01 |  |
 |
 |
| 1772-064-105_B02 |  |
 |
 |
| 1772-064-105_B03 |  |
 |
 |
| 1772-064-105_B04 |  |
 |
 |
| 1772-064-105_B05 |  |
 |
 |
| 1772-064-105_B06 |  |
 |
 |
| 1772-064-105_B07 |  |
 |
 |
| 1772-064-105_B08 |  |
 |
 |
| 1772-064-105_B09 |  |
 |
 |
| 1772-064-105_B10 |  |
 |
 |
| 1772-064-105_B11 |  |
 |
 |
| 1772-064-105_B12 |  |
 |
 |
| 1772-064-105_C01 |  |
 |
 |
| 1772-064-105_C02 |  |
 |
 |
| 1772-064-105_C03 |  |
 |
 |
| 1772-064-105_C04 |  |
 |
 |
| 1772-064-105_C05 |  |
 |
 |
| 1772-064-105_C06 |  |
 |
 |
| 1772-064-105_C07 |  |
 |
 |
| 1772-064-105_C08 |  |
 |
 |
| 1772-064-105_C09 |  |
 |
 |
| 1772-064-105_C10 |  |
 |
 |
| 1772-064-105_C11 |  |
 |
 |
| 1772-064-105_C12 |  |
 |
 |
| 1772-064-105_D01 |  |
 |
 |
| 1772-064-105_D02 |  |
 |
 |
| 1772-064-105_D03 |  |
 |
 |
| 1772-064-105_D04 |  |
 |
 |
| 1772-064-105_D05 |  |
 |
 |
| 1772-064-105_D06 |  |
 |
 |
| 1772-064-105_D07 |  |
 |
 |
| 1772-064-105_D08 |  |
 |
 |
| 1772-064-105_D09 |  |
 |
 |
| 1772-064-105_D10 |  |
 |
 |
| 1772-064-105_D11 |  |
 |
 |
| 1772-064-105_D12 |  |
 |
 |
| 1772-064-105_E01 |  |
 |
 |
| 1772-064-105_E02 |  |
 |
 |
| 1772-064-105_E03 |  |
 |
 |
| 1772-064-105_E04 |  |
 |
 |
| 1772-064-105_E05 |  |
 |
 |
| 1772-064-105_E06 |  |
 |
 |
| 1772-064-105_E07 |  |
 |
 |
| 1772-064-105_E08 |  |
 |
 |
| 1772-064-105_E09 |  |
 |
 |
| 1772-064-105_E10 |  |
 |
 |
| 1772-064-105_E11 |  |
 |
 |
| 1772-064-105_E12 |  |
 |
 |
| 1772-064-105_F01 |  |
 |
 |
| 1772-064-105_F02 |  |
 |
 |
| 1772-064-105_F03 |  |
 |
 |
| 1772-064-105_F04 |  |
 |
 |
| 1772-064-105_F05 |  |
 |
 |
| 1772-064-105_F06 |  |
 |
 |
| 1772-064-105_F07 |  |
 |
 |
| 1772-064-105_F08 |  |
 |
 |
| 1772-064-105_F09 |  |
 |
 |
| 1772-064-105_F10 |  |
 |
 |
| 1772-064-105_F11 |  |
 |
 |
| 1772-064-105_F12 |  |
 |
 |
| 1772-064-105_G01 |  |
 |
 |
| 1772-064-105_G02 |  |
 |
 |
| 1772-064-105_G03 |  |
 |
 |
| 1772-064-105_G04 |  |
 |
 |
| 1772-064-105_G05 |  |
 |
 |
| 1772-064-105_G06 |  |
 |
 |
| 1772-064-105_G07 |  |
 |
 |
| 1772-064-105_G08 |  |
 |
 |
| 1772-064-105_G09 |  |
 |
 |
| 1772-064-105_G10 |  |
 |
 |
| 1772-064-105_G11 |  |
 |
 |
| 1772-064-105_G12 |  |
 |
 |
| 1772-064-105_H01 |  |
 |
 |
| 1772-064-105_H02 |  |
 |
 |
| 1772-064-105_H03 |  |
 |
 |
| 1772-064-105_H04 |  |
 |
 |
| 1772-064-105_H05 |  |
 |
 |
| 1772-064-105_H06 |  |
 |
 |
| 1772-064-105_H07 |  |
 |
 |
| 1772-064-105_H08 |  |
 |
 |
| 1772-064-105_H09 |  |
 |
 |
| 1772-064-105_H10 |  |
 |
 |
| 1772-064-105_H11 |  |
 |
 |
| 1772-064-105_H12 |  |
 |
 |
| 1772-064-108_A01 |  |
 |
 |
| 1772-064-108_A02 |  |
 |
 |
| 1772-064-108_A03 |  |
 |
 |
| 1772-064-108_A04 |  |
 |
 |
| 1772-064-108_A05 |  |
 |
 |
| 1772-064-108_A06 |  |
 |
 |
| 1772-064-108_A07 |  |
 |
 |
| 1772-064-108_A08 |  |
 |
 |
| 1772-064-108_A09 |  |
 |
 |
| 1772-064-108_A10 |  |
 |
 |
| 1772-064-108_A11 |  |
 |
 |
| 1772-064-108_A12 |  |
 |
 |
| 1772-064-108_B01 |  |
 |
 |
| 1772-064-108_B02 |  |
 |
 |
| 1772-064-108_B03 |  |
 |
 |
| 1772-064-108_B04 |  |
 |
 |
| 1772-064-108_B05 |  |
 |
 |
| 1772-064-108_B06 |  |
 |
 |
| 1772-064-108_B07 |  |
 |
 |
| 1772-064-108_B08 |  |
 |
 |
| 1772-064-108_B09 |  |
 |
 |
| 1772-064-108_B10 |  |
 |
 |
| 1772-064-108_B11 |  |
 |
 |
| 1772-064-108_B12 |  |
 |
 |
| 1772-064-108_C01 |  |
 |
 |
| 1772-064-108_C02 |  |
 |
 |
| 1772-064-108_C03 |  |
 |
 |
| 1772-064-108_C04 |  |
 |
 |
| 1772-064-108_C05 |  |
 |
 |
| 1772-064-108_C06 |  |
 |
 |
| 1772-064-108_C07 |  |
 |
 |
| 1772-064-108_C08 |  |
 |
 |
| 1772-064-108_C09 |  |
 |
 |
| 1772-064-108_C10 |  |
 |
 |
| 1772-064-108_C11 |  |
 |
 |
| 1772-064-108_C12 |  |
 |
 |
| 1772-064-108_D01 |  |
 |
 |
| 1772-064-108_D02 |  |
 |
 |
| 1772-064-108_D03 |  |
 |
 |
| 1772-064-108_D04 |  |
 |
 |
| 1772-064-108_D05 |  |
 |
 |
| 1772-064-108_D06 |  |
 |
 |
| 1772-064-108_D07 |  |
 |
 |
| 1772-064-108_D08 |  |
 |
 |
| 1772-064-108_D09 |  |
 |
 |
| 1772-064-108_D10 |  |
 |
 |
| 1772-064-108_D11 |  |
 |
 |
| 1772-064-108_D12 |  |
 |
 |
| 1772-064-108_E01 |  |
 |
 |
| 1772-064-108_E02 |  |
 |
 |
| 1772-064-108_E03 |  |
 |
 |
| 1772-064-108_E04 |  |
 |
 |
| 1772-064-108_E05 |  |
 |
 |
| 1772-064-108_E06 |  |
 |
 |
| 1772-064-108_E07 |  |
 |
 |
| 1772-064-108_E08 |  |
 |
 |
| 1772-064-108_E09 |  |
 |
 |
| 1772-064-108_E10 |  |
 |
 |
| 1772-064-108_E11 |  |
 |
 |
| 1772-064-108_E12 |  |
 |
 |
| 1772-064-108_F01 |  |
 |
 |
| 1772-064-108_F02 |  |
 |
 |
| 1772-064-108_F03 |  |
 |
 |
| 1772-064-108_F04 |  |
 |
 |
| 1772-064-108_F05 |  |
 |
 |
| 1772-064-108_F06 |  |
 |
 |
| 1772-064-108_F07 |  |
 |
 |
| 1772-064-108_F08 |  |
 |
 |
| 1772-064-108_F09 |  |
 |
 |
| 1772-064-108_F10 |  |
 |
 |
| 1772-064-108_F11 |  |
 |
 |
| 1772-064-108_F12 |  |
 |
 |
| 1772-064-108_G01 |  |
 |
 |
| 1772-064-108_G02 |  |
 |
 |
| 1772-064-108_G03 |  |
 |
 |
| 1772-064-108_G04 |  |
 |
 |
| 1772-064-108_G05 |  |
 |
 |
| 1772-064-108_G06 |  |
 |
 |
| 1772-064-108_G07 |  |
 |
 |
| 1772-064-108_G08 |  |
 |
 |
| 1772-064-108_G09 |  |
 |
 |
| 1772-064-108_G10 |  |
 |
 |
| 1772-064-108_G11 |  |
 |
 |
| 1772-064-108_G12 |  |
 |
 |
| 1772-064-108_H01 |  |
 |
 |
| 1772-064-108_H02 |  |
 |
 |
| 1772-064-108_H03 |  |
 |
 |
| 1772-064-108_H04 |  |
 |
 |
| 1772-064-108_H05 |  |
 |
 |
| 1772-064-108_H06 |  |
 |
 |
| 1772-064-108_H07 |  |
 |
 |
| 1772-064-108_H08 |  |
 |
 |
| 1772-064-108_H09 |  |
 |
 |
| 1772-064-108_H10 |  |
 |
 |
| 1772-064-108_H11 |  |
 |
 |
| 1772-064-108_H12 |  |
 |
 |
| 1772-064-109_A01 |  |
 |
 |
| 1772-064-109_A02 |  |
 |
 |
| 1772-064-109_A03 |  |
 |
 |
| 1772-064-109_A04 |  |
 |
 |
| 1772-064-109_A05 |  |
 |
 |
| 1772-064-109_A06 |  |
 |
 |
| 1772-064-109_A07 |  |
 |
 |
| 1772-064-109_A08 |  |
 |
 |
| 1772-064-109_A09 |  |
 |
 |
| 1772-064-109_A10 |  |
 |
 |
| 1772-064-109_A11 |  |
 |
 |
| 1772-064-109_A12 |  |
 |
 |
| 1772-064-109_B01 |  |
 |
 |
| 1772-064-109_B02 |  |
 |
 |
| 1772-064-109_B03 |  |
 |
 |
| 1772-064-109_B04 |  |
 |
 |
| 1772-064-109_B05 |  |
 |
 |
| 1772-064-109_B06 |  |
 |
 |
| 1772-064-109_B07 |  |
 |
 |
| 1772-064-109_B08 |  |
 |
 |
| 1772-064-109_B09 |  |
 |
 |
| 1772-064-109_B10 |  |
 |
 |
| 1772-064-109_B11 |  |
 |
 |
| 1772-064-109_B12 |  |
 |
 |
| 1772-064-109_C01 |  |
 |
 |
| 1772-064-109_C02 |  |
 |
 |
| 1772-064-109_C03 |  |
 |
 |
| 1772-064-109_C04 |  |
 |
 |
| 1772-064-109_C05 |  |
 |
 |
| 1772-064-109_C06 |  |
 |
 |
| 1772-064-109_C07 |  |
 |
 |
| 1772-064-109_C08 |  |
 |
 |
| 1772-064-109_C09 |  |
 |
 |
| 1772-064-109_C10 |  |
 |
 |
| 1772-064-109_C11 |  |
 |
 |
| 1772-064-109_C12 |  |
 |
 |
| 1772-064-109_D01 |  |
 |
 |
| 1772-064-109_D02 |  |
 |
 |
| 1772-064-109_D03 |  |
 |
 |
| 1772-064-109_D04 |  |
 |
 |
| 1772-064-109_D05 |  |
 |
 |
| 1772-064-109_D06 |  |
 |
 |
| 1772-064-109_D07 |  |
 |
 |
| 1772-064-109_D08 |  |
 |
 |
| 1772-064-109_D09 |  |
 |
 |
| 1772-064-109_D10 |  |
 |
 |
| 1772-064-109_D11 |  |
 |
 |
| 1772-064-109_D12 |  |
 |
 |
| 1772-064-109_E01 |  |
 |
 |
| 1772-064-109_E02 |  |
 |
 |
| 1772-064-109_E03 |  |
 |
 |
| 1772-064-109_E04 |  |
 |
 |
| 1772-064-109_E05 |  |
 |
 |
| 1772-064-109_E06 |  |
 |
 |
| 1772-064-109_E07 |  |
 |
 |
| 1772-064-109_E08 |  |
 |
 |
| 1772-064-109_E09 |  |
 |
 |
| 1772-064-109_E10 |  |
 |
 |
| 1772-064-109_E11 |  |
 |
 |
| 1772-064-109_E12 |  |
 |
 |
| 1772-064-109_F01 |  |
 |
 |
| 1772-064-109_F02 |  |
 |
 |
| 1772-064-109_F03 |  |
 |
 |
| 1772-064-109_F04 |  |
 |
 |
| 1772-064-109_F05 |  |
 |
 |
| 1772-064-109_F06 |  |
 |
 |
| 1772-064-109_F07 |  |
 |
 |
| 1772-064-109_F08 |  |
 |
 |
| 1772-064-109_F09 |  |
 |
 |
| 1772-064-109_F10 |  |
 |
 |
| 1772-064-109_F11 |  |
 |
 |
| 1772-064-109_F12 |  |
 |
 |
| 1772-064-109_G01 |  |
 |
 |
| 1772-064-109_G02 |  |
 |
 |
| 1772-064-109_G03 |  |
 |
 |
| 1772-064-109_G04 |  |
 |
 |
| 1772-064-109_G05 |  |
 |
 |
| 1772-064-109_G06 |  |
 |
 |
| 1772-064-109_G07 |  |
 |
 |
| 1772-064-109_G08 |  |
 |
 |
| 1772-064-109_G09 |  |
 |
 |
| 1772-064-109_G10 |  |
 |
 |
| 1772-064-109_G11 |  |
 |
 |
| 1772-064-109_G12 |  |
 |
 |
| 1772-064-109_H01 |  |
 |
 |
| 1772-064-109_H02 |  |
 |
 |
| 1772-064-109_H03 |  |
 |
 |
| 1772-064-109_H04 |  |
 |
 |
| 1772-064-109_H05 |  |
 |
 |
| 1772-064-109_H06 |  |
 |
 |
| 1772-064-109_H07 |  |
 |
 |
| 1772-064-109_H08 |  |
 |
 |
| 1772-064-109_H09 |  |
 |
 |
| 1772-064-109_H10 |  |
 |
 |
| 1772-064-109_H11 |  |
 |
 |
| 1772-064-109_H12 |  |
 |
 |
| 1772-066-262_A01 |  |
 |
 |
| 1772-066-262_A02 |  |
 |
 |
| 1772-066-262_A03 |  |
 |
 |
| 1772-066-262_A04 |  |
 |
 |
| 1772-066-262_A05 |  |
 |
 |
| 1772-066-262_A06 |  |
 |
 |
| 1772-066-262_A07 |  |
 |
 |
| 1772-066-262_A08 |  |
 |
 |
| 1772-066-262_A09 |  |
 |
 |
| 1772-066-262_A10 |  |
 |
 |
| 1772-066-262_A11 |  |
 |
 |
| 1772-066-262_A12 |  |
 |
 |
| 1772-066-262_B01 |  |
 |
 |
| 1772-066-262_B02 |  |
 |
 |
| 1772-066-262_B03 |  |
 |
 |
| 1772-066-262_B04 |  |
 |
 |
| 1772-066-262_B05 |  |
 |
 |
| 1772-066-262_B06 |  |
 |
 |
| 1772-066-262_B07 |  |
 |
 |
| 1772-066-262_B08 |  |
 |
 |
| 1772-066-262_B09 |  |
 |
 |
| 1772-066-262_B10 |  |
 |
 |
| 1772-066-262_B11 |  |
 |
 |
| 1772-066-262_B12 |  |
 |
 |
| 1772-066-262_C01 |  |
 |
 |
| 1772-066-262_C02 |  |
 |
 |
| 1772-066-262_C03 |  |
 |
 |
| 1772-066-262_C04 |  |
 |
 |
| 1772-066-262_C05 |  |
 |
 |
| 1772-066-262_C06 |  |
 |
 |
| 1772-066-262_C07 |  |
 |
 |
| 1772-066-262_C08 |  |
 |
 |
| 1772-066-262_C09 |  |
 |
 |
| 1772-066-262_C10 |  |
 |
 |
| 1772-066-262_C11 |  |
 |
 |
| 1772-066-262_C12 |  |
 |
 |
| 1772-066-262_D01 |  |
 |
 |
| 1772-066-262_D02 |  |
 |
 |
| 1772-066-262_D03 |  |
 |
 |
| 1772-066-262_D04 |  |
 |
 |
| 1772-066-262_D05 |  |
 |
 |
| 1772-066-262_D06 |  |
 |
 |
| 1772-066-262_D07 |  |
 |
 |
| 1772-066-262_D08 |  |
 |
 |
| 1772-066-262_D09 |  |
 |
 |
| 1772-066-262_D10 |  |
 |
 |
| 1772-066-262_D11 |  |
 |
 |
| 1772-066-262_D12 |  |
 |
 |
| 1772-066-262_E01 |  |
 |
 |
| 1772-066-262_E02 |  |
 |
 |
| 1772-066-262_E03 |  |
 |
 |
| 1772-066-262_E04 |  |
 |
 |
| 1772-066-262_E05 |  |
 |
 |
| 1772-066-262_E06 |  |
 |
 |
| 1772-066-262_E07 |  |
 |
 |
| 1772-066-262_E08 |  |
 |
 |
| 1772-066-262_E09 |  |
 |
 |
| 1772-066-262_E10 |  |
 |
 |
| 1772-066-262_E11 |  |
 |
 |
| 1772-066-262_E12 |  |
 |
 |
| 1772-066-262_F01 |  |
 |
 |
| 1772-066-262_F02 |  |
 |
 |
| 1772-066-262_F03 |  |
 |
 |
| 1772-066-262_F04 |  |
 |
 |
| 1772-066-262_F05 |  |
 |
 |
| 1772-066-262_F06 |  |
 |
 |
| 1772-066-262_F07 |  |
 |
 |
| 1772-066-262_F08 |  |
 |
 |
| 1772-066-262_F09 |  |
 |
 |
| 1772-066-262_F10 |  |
 |
 |
| 1772-066-262_F11 |  |
 |
 |
| 1772-066-262_F12 |  |
 |
 |
| 1772-066-262_G01 |  |
 |
 |
| 1772-066-262_G02 |  |
 |
 |
| 1772-066-262_G03 |  |
 |
 |
| 1772-066-262_G04 |  |
 |
 |
| 1772-066-262_G05 |  |
 |
 |
| 1772-066-262_G06 |  |
 |
 |
| 1772-066-262_G07 |  |
 |
 |
| 1772-066-262_G08 |  |
 |
 |
| 1772-066-262_G09 |  |
 |
 |
| 1772-066-262_G10 |  |
 |
 |
| 1772-066-262_G11 |  |
 |
 |
| 1772-066-262_G12 |  |
 |
 |
| 1772-066-262_H01 |  |
 |
 |
| 1772-066-262_H02 |  |
 |
 |
| 1772-066-262_H03 |  |
 |
 |
| 1772-066-262_H04 |  |
 |
 |
| 1772-066-262_H05 |  |
 |
 |
| 1772-066-262_H06 |  |
 |
 |
| 1772-066-262_H07 |  |
 |
 |
| 1772-066-262_H08 |  |
 |
 |
| 1772-066-262_H09 |  |
 |
 |
| 1772-066-262_H10 |  |
 |
 |
| 1772-066-262_H11 |  |
 |
 |
| 1772-066-262_H12 |  |
 |
 |
| 1772-066-263_A01 |  |
 |
 |
| 1772-066-263_A02 |  |
 |
 |
| 1772-066-263_A03 |  |
 |
 |
| 1772-066-263_A04 |  |
 |
 |
| 1772-066-263_A05 |  |
 |
 |
| 1772-066-263_A06 |  |
 |
 |
| 1772-066-263_A07 |  |
 |
 |
| 1772-066-263_A08 |  |
 |
 |
| 1772-066-263_A09 |  |
 |
 |
| 1772-066-263_A10 |  |
 |
 |
| 1772-066-263_A11 |  |
 |
 |
| 1772-066-263_A12 |  |
 |
 |
| 1772-066-263_B01 |  |
 |
 |
| 1772-066-263_B02 |  |
 |
 |
| 1772-066-263_B03 |  |
 |
 |
| 1772-066-263_B04 |  |
 |
 |
| 1772-066-263_B05 |  |
 |
 |
| 1772-066-263_B06 |  |
 |
 |
| 1772-066-263_B07 |  |
 |
 |
| 1772-066-263_B08 |  |
 |
 |
| 1772-066-263_B09 |  |
 |
 |
| 1772-066-263_B10 |  |
 |
 |
| 1772-066-263_B11 |  |
 |
 |
| 1772-066-263_B12 |  |
 |
 |
| 1772-066-263_C01 |  |
 |
 |
| 1772-066-263_C02 |  |
 |
 |
| 1772-066-263_C03 |  |
 |
 |
| 1772-066-263_C04 |  |
 |
 |
| 1772-066-263_C05 |  |
 |
 |
| 1772-066-263_C06 |  |
 |
 |
| 1772-066-263_C07 |  |
 |
 |
| 1772-066-263_C08 |  |
 |
 |
| 1772-066-263_C09 |  |
 |
 |
| 1772-066-263_C10 |  |
 |
 |
| 1772-066-263_C11 |  |
 |
 |
| 1772-066-263_C12 |  |
 |
 |
| 1772-066-263_D01 |  |
 |
 |
| 1772-066-263_D02 |  |
 |
 |
| 1772-066-263_D03 |  |
 |
 |
| 1772-066-263_D04 |  |
 |
 |
| 1772-066-263_D05 |  |
 |
 |
| 1772-066-263_D06 |  |
 |
 |
| 1772-066-263_D07 |  |
 |
 |
| 1772-066-263_D08 |  |
 |
 |
| 1772-066-263_D09 |  |
 |
 |
| 1772-066-263_D10 |  |
 |
 |
| 1772-066-263_D11 |  |
 |
 |
| 1772-066-263_D12 |  |
 |
 |
| 1772-066-263_E01 |  |
 |
 |
| 1772-066-263_E02 |  |
 |
 |
| 1772-066-263_E03 |  |
 |
 |
| 1772-066-263_E04 |  |
 |
 |
| 1772-066-263_E05 |  |
 |
 |
| 1772-066-263_E06 |  |
 |
 |
| 1772-066-263_E07 |  |
 |
 |
| 1772-066-263_E08 |  |
 |
 |
| 1772-066-263_E09 |  |
 |
 |
| 1772-066-263_E10 |  |
 |
 |
| 1772-066-263_E11 |  |
 |
 |
| 1772-066-263_E12 |  |
 |
 |
| 1772-066-263_F01 |  |
 |
 |
| 1772-066-263_F02 |  |
 |
 |
| 1772-066-263_F03 |  |
 |
 |
| 1772-066-263_F04 |  |
 |
 |
| 1772-066-263_F05 |  |
 |
 |
| 1772-066-263_F06 |  |
 |
 |
| 1772-066-263_F07 |  |
 |
 |
| 1772-066-263_F08 |  |
 |
 |
| 1772-066-263_F09 |  |
 |
 |
| 1772-066-263_F10 |  |
 |
 |
| 1772-066-263_F11 |  |
 |
 |
| 1772-066-263_F12 |  |
 |
 |
| 1772-066-263_G01 |  |
 |
 |
| 1772-066-263_G02 |  |
 |
 |
| 1772-066-263_G03 |  |
 |
 |
| 1772-066-263_G04 |  |
 |
 |
| 1772-066-263_G05 |  |
 |
 |
| 1772-066-263_G06 |  |
 |
 |
| 1772-066-263_G07 |  |
 |
 |
| 1772-066-263_G08 |  |
 |
 |
| 1772-066-263_G09 |  |
 |
 |
| 1772-066-263_G10 |  |
 |
 |
| 1772-066-263_G11 |  |
 |
 |
| 1772-066-263_G12 |  |
 |
 |
| 1772-066-263_H01 |  |
 |
 |
| 1772-066-263_H02 |  |
 |
 |
| 1772-066-263_H03 |  |
 |
 |
| 1772-066-263_H04 |  |
 |
 |
| 1772-066-263_H05 |  |
 |
 |
| 1772-066-263_H06 |  |
 |
 |
| 1772-066-263_H07 |  |
 |
 |
| 1772-066-263_H08 |  |
 |
 |
| 1772-066-263_H09 |  |
 |
 |
| 1772-066-263_H10 |  |
 |
 |
| 1772-066-263_H11 |  |
 |
 |
| 1772-066-263_H12 |  |
 |
 |
| 1772-067-035_A01 |  |
 |
 |
| 1772-067-035_A02 |  |
 |
 |
| 1772-067-035_A03 |  |
 |
 |
| 1772-067-035_A04 |  |
 |
 |
| 1772-067-035_A05 |  |
 |
 |
| 1772-067-035_A06 |  |
 |
 |
| 1772-067-035_A07 |  |
 |
 |
| 1772-067-035_A08 |  |
 |
 |
| 1772-067-035_A09 |  |
 |
 |
| 1772-067-035_A10 |  |
 |
 |
| 1772-067-035_A11 |  |
 |
 |
| 1772-067-035_A12 |  |
 |
 |
| 1772-067-035_B01 |  |
 |
 |
| 1772-067-035_B02 |  |
 |
 |
| 1772-067-035_B03 |  |
 |
 |
| 1772-067-035_B04 |  |
 |
 |
| 1772-067-035_B05 |  |
 |
 |
| 1772-067-035_B06 |  |
 |
 |
| 1772-067-035_B07 |  |
 |
 |
| 1772-067-035_B08 |  |
 |
 |
| 1772-067-035_B09 |  |
 |
 |
| 1772-067-035_B10 |  |
 |
 |
| 1772-067-035_B11 |  |
 |
 |
| 1772-067-035_B12 |  |
 |
 |
| 1772-067-035_C01 |  |
 |
 |
| 1772-067-035_C02 |  |
 |
 |
| 1772-067-035_C03 |  |
 |
 |
| 1772-067-035_C04 |  |
 |
 |
| 1772-067-035_C05 |  |
 |
 |
| 1772-067-035_C06 |  |
 |
 |
| 1772-067-035_C07 |  |
 |
 |
| 1772-067-035_C08 |  |
 |
 |
| 1772-067-035_C09 |  |
 |
 |
| 1772-067-035_C10 |  |
 |
 |
| 1772-067-035_C11 |  |
 |
 |
| 1772-067-035_C12 |  |
 |
 |
| 1772-067-035_D01 |  |
 |
 |
| 1772-067-035_D02 |  |
 |
 |
| 1772-067-035_D03 |  |
 |
 |
| 1772-067-035_D04 |  |
 |
 |
| 1772-067-035_D05 |  |
 |
 |
| 1772-067-035_D06 |  |
 |
 |
| 1772-067-035_D07 |  |
 |
 |
| 1772-067-035_D08 |  |
 |
 |
| 1772-067-035_D09 |  |
 |
 |
| 1772-067-035_D10 |  |
 |
 |
| 1772-067-035_D11 |  |
 |
 |
| 1772-067-035_D12 |  |
 |
 |
| 1772-067-035_E01 |  |
 |
 |
| 1772-067-035_E02 |  |
 |
 |
| 1772-067-035_E03 |  |
 |
 |
| 1772-067-035_E04 |  |
 |
 |
| 1772-067-035_E05 |  |
 |
 |
| 1772-067-035_E06 |  |
 |
 |
| 1772-067-035_E07 |  |
 |
 |
| 1772-067-035_E08 |  |
 |
 |
| 1772-067-035_E09 |  |
 |
 |
| 1772-067-035_E10 |  |
 |
 |
| 1772-067-035_E11 |  |
 |
 |
| 1772-067-035_E12 |  |
 |
 |
| 1772-067-035_F01 |  |
 |
 |
| 1772-067-035_F02 |  |
 |
 |
| 1772-067-035_F03 |  |
 |
 |
| 1772-067-035_F04 |  |
 |
 |
| 1772-067-035_F05 |  |
 |
 |
| 1772-067-035_F06 |  |
 |
 |
| 1772-067-035_F07 |  |
 |
 |
| 1772-067-035_F08 |  |
 |
 |
| 1772-067-035_F09 |  |
 |
 |
| 1772-067-035_F10 |  |
 |
 |
| 1772-067-035_F11 |  |
 |
 |
| 1772-067-035_F12 |  |
 |
 |
| 1772-067-035_G01 |  |
 |
 |
| 1772-067-035_G02 |  |
 |
 |
| 1772-067-035_G03 |  |
 |
 |
| 1772-067-035_G04 |  |
 |
 |
| 1772-067-035_G05 |  |
 |
 |
| 1772-067-035_G06 |  |
 |
 |
| 1772-067-035_G07 |  |
 |
 |
| 1772-067-035_G08 |  |
 |
 |
| 1772-067-035_G09 |  |
 |
 |
| 1772-067-035_G10 |  |
 |
 |
| 1772-067-035_G11 |  |
 |
 |
| 1772-067-035_G12 |  |
 |
 |
| 1772-067-035_H01 |  |
 |
 |
| 1772-067-035_H02 |  |
 |
 |
| 1772-067-035_H03 |  |
 |
 |
| 1772-067-035_H04 |  |
 |
 |
| 1772-067-035_H05 |  |
 |
 |
| 1772-067-035_H06 |  |
 |
 |
| 1772-067-035_H07 |  |
 |
 |
| 1772-067-035_H08 |  |
 |
 |
| 1772-067-035_H09 |  |
 |
 |
| 1772-067-035_H10 |  |
 |
 |
| 1772-067-035_H11 |  |
 |
 |
| 1772-067-035_H12 |  |
 |
 |
| 1772-115-303_A01 |  |
 |
 |
| 1772-115-303_A02 |  |
 |
 |
| 1772-115-303_A03 |  |
 |
 |
| 1772-115-303_A04 |  |
 |
 |
| 1772-115-303_A05 |  |
 |
 |
| 1772-115-303_A06 |  |
 |
 |
| 1772-115-303_A07 |  |
 |
 |
| 1772-115-303_A08 |  |
 |
 |
| 1772-115-303_A09 |  |
 |
 |
| 1772-115-303_A10 |  |
 |
 |
| 1772-115-303_A11 |  |
 |
 |
| 1772-115-303_A12 |  |
 |
 |
| 1772-115-303_B01 |  |
 |
 |
| 1772-115-303_B02 |  |
 |
 |
| 1772-115-303_B03 |  |
 |
 |
| 1772-115-303_B04 |  |
 |
 |
| 1772-115-303_B05 |  |
 |
 |
| 1772-115-303_B06 |  |
 |
 |
| 1772-115-303_B07 |  |
 |
 |
| 1772-115-303_B08 |  |
 |
 |
| 1772-115-303_B09 |  |
 |
 |
| 1772-115-303_B10 |  |
 |
 |
| 1772-115-303_B11 |  |
 |
 |
| 1772-115-303_B12 |  |
 |
 |
| 1772-115-303_C01 |  |
 |
 |
| 1772-115-303_C02 |  |
 |
 |
| 1772-115-303_C03 |  |
 |
 |
| 1772-115-303_C04 |  |
 |
 |
| 1772-115-303_C05 |  |
 |
 |
| 1772-115-303_C06 |  |
 |
 |
| 1772-115-303_C07 |  |
 |
 |
| 1772-115-303_C08 |  |
 |
 |
| 1772-115-303_C09 |  |
 |
 |
| 1772-115-303_C10 |  |
 |
 |
| 1772-115-303_C11 |  |
 |
 |
| 1772-115-303_C12 |  |
 |
 |
| 1772-115-303_D01 |  |
 |
 |
| 1772-115-303_D02 |  |
 |
 |
| 1772-115-303_D03 |  |
 |
 |
| 1772-115-303_D04 |  |
 |
 |
| 1772-115-303_D05 |  |
 |
 |
| 1772-115-303_D06 |  |
 |
 |
| 1772-115-303_D07 |  |
 |
 |
| 1772-115-303_D08 |  |
 |
 |
| 1772-115-303_D09 |  |
 |
 |
| 1772-115-303_D10 |  |
 |
 |
| 1772-115-303_D11 |  |
 |
 |
| 1772-115-303_D12 |  |
 |
 |
| 1772-115-303_E01 |  |
 |
 |
| 1772-115-303_E02 |  |
 |
 |
| 1772-115-303_E03 |  |
 |
 |
| 1772-115-303_E04 |  |
 |
 |
| 1772-115-303_E05 |  |
 |
 |
| 1772-115-303_E06 |  |
 |
 |
| 1772-115-303_E07 |  |
 |
 |
| 1772-115-303_E08 |  |
 |
 |
| 1772-115-303_E09 |  |
 |
 |
| 1772-115-303_E10 |  |
 |
 |
| 1772-115-303_E11 |  |
 |
 |
| 1772-115-303_E12 |  |
 |
 |
| 1772-115-303_F01 |  |
 |
 |
| 1772-115-303_F02 |  |
 |
 |
| 1772-115-303_F03 |  |
 |
 |
| 1772-115-303_F04 |  |
 |
 |
| 1772-115-303_F05 |  |
 |
 |
| 1772-115-303_F06 |  |
 |
 |
| 1772-115-303_F07 |  |
 |
 |
| 1772-115-303_F08 |  |
 |
 |
| 1772-115-303_F09 |  |
 |
 |
| 1772-115-303_F10 |  |
 |
 |
| 1772-115-303_F11 |  |
 |
 |
| 1772-115-303_F12 |  |
 |
 |
| 1772-115-303_G01 |  |
 |
 |
| 1772-115-303_G02 |  |
 |
 |
| 1772-115-303_G03 |  |
 |
 |
| 1772-115-303_G04 |  |
 |
 |
| 1772-115-303_G05 |  |
 |
 |
| 1772-115-303_G06 |  |
 |
 |
| 1772-115-303_G07 |  |
 |
 |
| 1772-115-303_G08 |  |
 |
 |
| 1772-115-303_G09 |  |
 |
 |
| 1772-115-303_G10 |  |
 |
 |
| 1772-115-303_G11 |  |
 |
 |
| 1772-115-303_G12 |  |
 |
 |
| 1772-115-303_H01 |  |
 |
 |
| 1772-115-303_H02 |  |
 |
 |
| 1772-115-303_H03 |  |
 |
 |
| 1772-115-303_H04 |  |
 |
 |
| 1772-115-303_H05 |  |
 |
 |
| 1772-115-303_H06 |  |
 |
 |
| 1772-115-303_H07 |  |
 |
 |
| 1772-115-303_H08 |  |
 |
 |
| 1772-115-303_H09 |  |
 |
 |
| 1772-115-303_H10 |  |
 |
 |
| 1772-115-303_H11 |  |
 |
 |
| 1772-115-303_H12 |  |
 |
 |
| 1772-115-305_A01 |  |
 |
 |
| 1772-115-305_A02 |  |
 |
 |
| 1772-115-305_A03 |  |
 |
 |
| 1772-115-305_A04 |  |
 |
 |
| 1772-115-305_A05 |  |
 |
 |
| 1772-115-305_A06 |  |
 |
 |
| 1772-115-305_A07 |  |
 |
 |
| 1772-115-305_A08 |  |
 |
 |
| 1772-115-305_A09 |  |
 |
 |
| 1772-115-305_A10 |  |
 |
 |
| 1772-115-305_A11 |  |
 |
 |
| 1772-115-305_A12 |  |
 |
 |
| 1772-115-305_B01 |  |
 |
 |
| 1772-115-305_B02 |  |
 |
 |
| 1772-115-305_B03 |  |
 |
 |
| 1772-115-305_B04 |  |
 |
 |
| 1772-115-305_B05 |  |
 |
 |
| 1772-115-305_B06 |  |
 |
 |
| 1772-115-305_B07 |  |
 |
 |
| 1772-115-305_B08 |  |
 |
 |
| 1772-115-305_B09 |  |
 |
 |
| 1772-115-305_B10 |  |
 |
 |
| 1772-115-305_B11 |  |
 |
 |
| 1772-115-305_B12 |  |
 |
 |
| 1772-115-305_C01 |  |
 |
 |
| 1772-115-305_C02 |  |
 |
 |
| 1772-115-305_C03 |  |
 |
 |
| 1772-115-305_C04 |  |
 |
 |
| 1772-115-305_C05 |  |
 |
 |
| 1772-115-305_C06 |  |
 |
 |
| 1772-115-305_C07 |  |
 |
 |
| 1772-115-305_C08 |  |
 |
 |
| 1772-115-305_C09 |  |
 |
 |
| 1772-115-305_C10 |  |
 |
 |
| 1772-115-305_C11 |  |
 |
 |
| 1772-115-305_C12 |  |
 |
 |
| 1772-115-305_D01 |  |
 |
 |
| 1772-115-305_D02 |  |
 |
 |
| 1772-115-305_D03 |  |
 |
 |
| 1772-115-305_D04 |  |
 |
 |
| 1772-115-305_D05 |  |
 |
 |
| 1772-115-305_D06 |  |
 |
 |
| 1772-115-305_D07 |  |
 |
 |
| 1772-115-305_D08 |  |
 |
 |
| 1772-115-305_D09 |  |
 |
 |
| 1772-115-305_D10 |  |
 |
 |
| 1772-115-305_D11 |  |
 |
 |
| 1772-115-305_D12 |  |
 |
 |
| 1772-115-305_E01 |  |
 |
 |
| 1772-115-305_E02 | 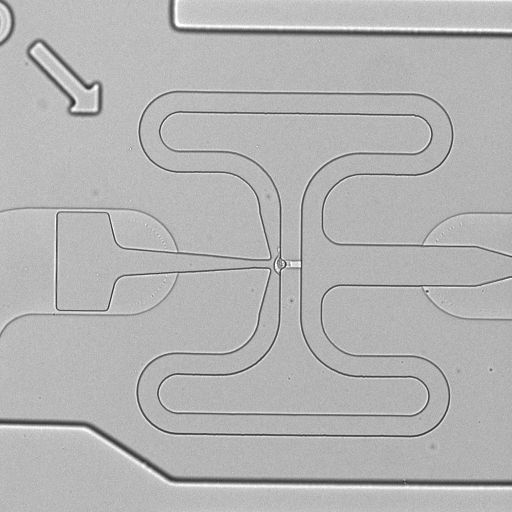 |
 |
 |
| 1772-115-305_E03 |  |
 |
 |
| 1772-115-305_E04 |  |
 |
 |
| 1772-115-305_E05 |  |
 |
 |
| 1772-115-305_E06 |  |
 |
 |
| 1772-115-305_E07 |  |
 |
 |
| 1772-115-305_E08 |  |
 |
 |
| 1772-115-305_E09 |  |
 |
 |
| 1772-115-305_E10 |  |
 |
 |
| 1772-115-305_E11 |  |
 |
 |
| 1772-115-305_E12 |  |
 |
 |
| 1772-115-305_F01 |  |
 |
 |
| 1772-115-305_F02 |  |
 |
 |
| 1772-115-305_F03 |  |
 |
 |
| 1772-115-305_F04 |  |
 |
 |
| 1772-115-305_F05 |  |
 |
 |
| 1772-115-305_F06 |  |
 |
 |
| 1772-115-305_F07 |  |
 |
 |
| 1772-115-305_F08 |  |
 |
 |
| 1772-115-305_F09 |  |
 |
 |
| 1772-115-305_F10 |  |
 |
 |
| 1772-115-305_F11 |  |
 |
 |
| 1772-115-305_F12 |  |
 |
 |
| 1772-115-305_G01 |  |
 |
 |
| 1772-115-305_G02 |  |
 |
 |
| 1772-115-305_G03 |  |
 |
 |
| 1772-115-305_G04 |  |
 |
 |
| 1772-115-305_G05 |  |
 |
 |
| 1772-115-305_G06 |  |
 |
 |
| 1772-115-305_G07 |  |
 |
 |
| 1772-115-305_G08 |  |
 |
 |
| 1772-115-305_G09 |  |
 |
 |
| 1772-115-305_G10 |  |
 |
 |
| 1772-115-305_G11 |  |
 |
 |
| 1772-115-305_G12 |  |
 |
 |
| 1772-115-305_H01 |  |
 |
 |
| 1772-115-305_H02 |  |
 |
 |
| 1772-115-305_H03 |  |
 |
 |
| 1772-115-305_H04 |  |
 |
 |
| 1772-115-305_H05 |  |
 |
 |
| 1772-115-305_H06 |  |
 |
 |
| 1772-115-305_H07 |  |
 |
 |
| 1772-115-305_H08 |  |
 |
 |
| 1772-115-305_H09 |  |
 |
 |
| 1772-115-305_H10 |  |
 |
 |
| 1772-115-305_H11 |  |
 |
 |
| 1772-115-305_H12 |  |
 |
 |
| 1772-115-308_A01 |  |
 |
 |
| 1772-115-308_A02 |  |
 |
 |
| 1772-115-308_A03 |  |
 |
 |
| 1772-115-308_A04 |  |
 |
 |
| 1772-115-308_A05 |  |
 |
 |
| 1772-115-308_A06 |  |
 |
 |
| 1772-115-308_A07 |  |
 |
 |
| 1772-115-308_A08 |  |
 |
 |
| 1772-115-308_A09 |  |
 |
 |
| 1772-115-308_A10 |  |
 |
 |
| 1772-115-308_A11 |  |
 |
 |
| 1772-115-308_A12 |  |
 |
 |
| 1772-115-308_B01 |  |
 |
 |
| 1772-115-308_B02 |  |
 |
 |
| 1772-115-308_B03 |  |
 |
 |
| 1772-115-308_B04 |  |
 |
 |
| 1772-115-308_B05 |  |
 |
 |
| 1772-115-308_B06 |  |
 |
 |
| 1772-115-308_B07 |  |
 |
 |
| 1772-115-308_B08 |  |
 |
 |
| 1772-115-308_B09 |  |
 |
 |
| 1772-115-308_B10 |  |
 |
 |
| 1772-115-308_B11 |  |
 |
 |
| 1772-115-308_B12 |  |
 |
 |
| 1772-115-308_C01 |  |
 |
 |
| 1772-115-308_C02 |  |
 |
 |
| 1772-115-308_C03 |  |
 |
 |
| 1772-115-308_C04 |  |
 |
 |
| 1772-115-308_C05 |  |
 |
 |
| 1772-115-308_C06 |  |
 |
 |
| 1772-115-308_C07 |  |
 |
 |
| 1772-115-308_C08 |  |
 |
 |
| 1772-115-308_C09 |  |
 |
 |
| 1772-115-308_C10 |  |
 |
 |
| 1772-115-308_C11 |  |
 |
 |
| 1772-115-308_C12 |  |
 |
 |
| 1772-115-308_D01 |  |
 |
 |
| 1772-115-308_D02 |  |
 |
 |
| 1772-115-308_D03 |  |
 |
 |
| 1772-115-308_D04 |  |
 |
 |
| 1772-115-308_D05 |  |
 |
 |
| 1772-115-308_D06 |  |
 |
 |
| 1772-115-308_D07 |  |
 |
 |
| 1772-115-308_D08 |  |
 |
 |
| 1772-115-308_D09 |  |
 |
 |
| 1772-115-308_D10 |  |
 |
 |
| 1772-115-308_D11 |  |
 |
 |
| 1772-115-308_D12 |  |
 |
 |
| 1772-115-308_E01 |  |
 |
 |
| 1772-115-308_E02 |  |
 |
 |
| 1772-115-308_E03 |  |
 |
 |
| 1772-115-308_E04 |  |
 |
 |
| 1772-115-308_E05 |  |
 |
 |
| 1772-115-308_E06 |  |
 |
 |
| 1772-115-308_E07 |  |
 |
 |
| 1772-115-308_E08 |  |
 |
 |
| 1772-115-308_E09 |  |
 |
 |
| 1772-115-308_E10 |  |
 |
 |
| 1772-115-308_E11 |  |
 |
 |
| 1772-115-308_E12 |  |
 |
 |
| 1772-115-308_F01 |  |
 |
 |
| 1772-115-308_F02 |  |
 |
 |
| 1772-115-308_F03 |  |
 |
 |
| 1772-115-308_F04 |  |
 |
 |
| 1772-115-308_F05 |  |
 |
 |
| 1772-115-308_F06 |  |
 |
 |
| 1772-115-308_F07 |  |
 |
 |
| 1772-115-308_F08 |  |
 |
 |
| 1772-115-308_F09 |  |
 |
 |
| 1772-115-308_F10 |  |
 |
 |
| 1772-115-308_F11 |  |
 |
 |
| 1772-115-308_F12 |  |
 |
 |
| 1772-115-308_G01 |  |
 |
 |
| 1772-115-308_G02 |  |
 |
 |
| 1772-115-308_G03 |  |
 |
 |
| 1772-115-308_G04 |  |
 |
 |
| 1772-115-308_G05 |  |
 |
 |
| 1772-115-308_G06 |  |
 |
 |
| 1772-115-308_G07 |  |
 |
 |
| 1772-115-308_G08 |  |
 |
 |
| 1772-115-308_G09 |  |
 |
 |
| 1772-115-308_G10 |  |
 |
 |
| 1772-115-308_G11 |  |
 |
 |
| 1772-115-308_G12 |  |
 |
 |
| 1772-115-308_H01 |  |
 |
 |
| 1772-115-308_H02 |  |
 |
 |
| 1772-115-308_H03 |  |
 |
 |
| 1772-115-308_H04 |  |
 |
 |
| 1772-115-308_H05 |  |
 |
 |
| 1772-115-308_H06 |  |
 |
 |
| 1772-115-308_H07 |  |
 |
 |
| 1772-115-308_H08 |  |
 |
 |
| 1772-115-308_H09 |  |
 |
 |
| 1772-115-308_H10 |  |
 |
 |
| 1772-115-308_H11 |  |
 |
 |
| 1772-115-308_H12 |  |
 |
 |
| 1772-115-310_A01 |  |
 |
 |
| 1772-115-310_A02 |  |
 |
 |
| 1772-115-310_A03 |  |
 |
 |
| 1772-115-310_A04 |  |
 |
 |
| 1772-115-310_A05 |  |
 |
 |
| 1772-115-310_A06 |  |
 |
 |
| 1772-115-310_A07 |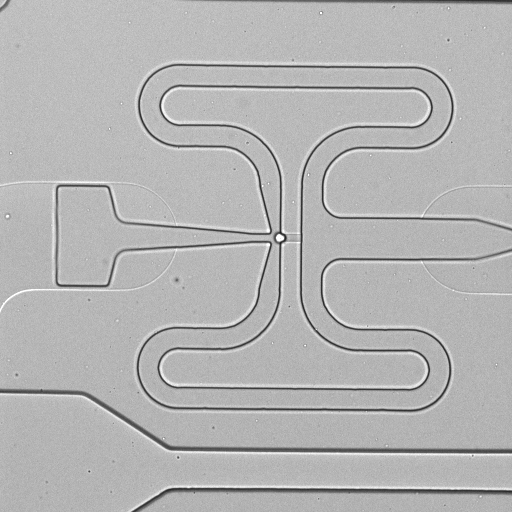  |
 |
 |
| 1772-115-310_A08 |  |
 |
 |
| 1772-115-310_A09 |  |
 |
 |
| 1772-115-310_A10 |  |
 |
 |
| 1772-115-310_A11 |  |
 |
 |
| 1772-115-310_A12 |  |
 |
 |
| 1772-115-310_B01 |  |
 |
 |
| 1772-115-310_B02 |  |
 |
 |
| 1772-115-310_B03 |  |
 |
 |
| 1772-115-310_B04 |  |
 |
 |
| 1772-115-310_B05 |  |
 |
 |
| 1772-115-310_B06 |  |
 |
 |
| 1772-115-310_B07 |  |
 |
 |
| 1772-115-310_B08 |  |
 |
 |
| 1772-115-310_B09 |  |
 |
 |
| 1772-115-310_B10 |  |
 |
 |
| 1772-115-310_B11 |  |
 |
 |
| 1772-115-310_B12 |  |
 |
 |
| 1772-115-310_C01 |  |
 |
 |
| 1772-115-310_C02 |  |
 |
 |
| 1772-115-310_C03 |  |
 |
 |
| 1772-115-310_C04 |  |
 |
 |
| 1772-115-310_C05 |  |
 |
 |
| 1772-115-310_C06 |  |
 |
 |
| 1772-115-310_C07 |  |
 |
 |
| 1772-115-310_C08 |  |
 |
 |
| 1772-115-310_C09 |  |
 |
 |
| 1772-115-310_C10 |  |
 |
 |
| 1772-115-310_C11 |  |
 |
 |
| 1772-115-310_C12 |  |
 |
 |
| 1772-115-310_D01 |  |
 |
 |
| 1772-115-310_D02 |  |
 |
 |
| 1772-115-310_D03 |  |
 |
 |
| 1772-115-310_D04 |  |
 |
 |
| 1772-115-310_D05 |  |
 |
 |
| 1772-115-310_D06 |  |
 |
 |
| 1772-115-310_D07 |  |
 |
 |
| 1772-115-310_D08 |  |
 |
 |
| 1772-115-310_D09 |  |
 |
 |
| 1772-115-310_D10 |  |
 |
 |
| 1772-115-310_D11 |  |
 |
 |
| 1772-115-310_D12 |  |
 |
 |
| 1772-115-310_E01 |  |
 |
 |
| 1772-115-310_E02 |  |
 |
 |
| 1772-115-310_E03 |  |
 |
 |
| 1772-115-310_E04 |  |
 |
 |
| 1772-115-310_E05 |  |
 |
 |
| 1772-115-310_E06 |  |
 |
 |
| 1772-115-310_E07 |  |
 |
 |
| 1772-115-310_E08 |  |
 |
 |
| 1772-115-310_E09 |  |
 |
 |
| 1772-115-310_E10 |  |
 |
 |
| 1772-115-310_E11 |  |
 |
 |
| 1772-115-310_E12 |  |
 |
 |
| 1772-115-310_F01 |  |
 |
 |
| 1772-115-310_F02 |  |
 |
 |
| 1772-115-310_F03 |  |
 |
 |
| 1772-115-310_F04 |  |
 |
 |
| 1772-115-310_F05 |  |
 |
 |
| 1772-115-310_F06 | 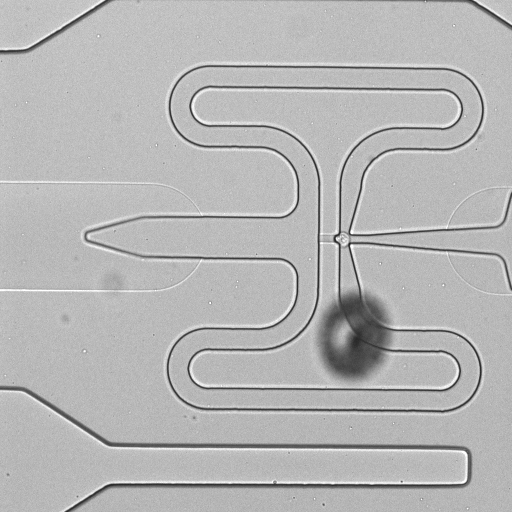 |
 |
 |
| 1772-115-310_F07 |  |
 |
 |
| 1772-115-310_F08 |  |
 |
 |
| 1772-115-310_F09 |  |
 |
 |
| 1772-115-310_F10 |  |
 |
 |
| 1772-115-310_F11 |  |
 |
 |
| 1772-115-310_F12 |  |
 |
 |
| 1772-115-310_G01 |  |
 |
 |
| 1772-115-310_G02 |  |
 |
 |
| 1772-115-310_G03 |  |
 |
 |
| 1772-115-310_G04 |  |
 |
 |
| 1772-115-310_G05 |  |
 |
 |
| 1772-115-310_G06 |  |
 |
 |
| 1772-115-310_G07 |  |
 |
 |
| 1772-115-310_G08 |  |
 |
 |
| 1772-115-310_G09 |  |
 |
 |
| 1772-115-310_G10 |  |
 |
 |
| 1772-115-310_G11 |  |
 |
 |
| 1772-115-310_G12 |  |
 |
 |
| 1772-115-310_H01 |  |
 |
 |
| 1772-115-310_H02 |  |
 |
 |
| 1772-115-310_H03 |  |
 |
 |
| 1772-115-310_H04 |  |
 |
 |
| 1772-115-310_H05 |  |
 |
 |
| 1772-115-310_H06 |  |
 |
 |
| 1772-115-310_H07 |  |
 |
 |
| 1772-115-310_H08 |  |
 |
 |
| 1772-115-310_H09 |  |
 |
 |
| 1772-115-310_H10 |  |
 |
 |
| 1772-115-310_H11 |  |
 |
 |
| 1772-115-310_H12 |  |
 |
 |
| 1772-115-311_A01 |  |
 |
 |
| 1772-115-311_A02 |  |
 |
 |
| 1772-115-311_A03 |  |
 |
 |
| 1772-115-311_A04 |  |
 |
 |
| 1772-115-311_A05 |  |
 |
 |
| 1772-115-311_A06 |  |
 |
 |
| 1772-115-311_A07 |  |
 |
 |
| 1772-115-311_A08 |  |
 |
 |
| 1772-115-311_A09 |  |
 |
 |
| 1772-115-311_A10 |  |
 |
 |
| 1772-115-311_A11 |  |
 |
 |
| 1772-115-311_A12 |  |
 |
 |
| 1772-115-311_B01 |  |
 |
 |
| 1772-115-311_B02 |  |
 |
 |
| 1772-115-311_B03 |  |
 |
 |
| 1772-115-311_B04 |  |
 |
 |
| 1772-115-311_B05 |  |
 |
 |
| 1772-115-311_B06 |  |
 |
 |
| 1772-115-311_B07 |  |
 |
 |
| 1772-115-311_B08 |  |
 |
 |
| 1772-115-311_B09 |  |
 |
 |
| 1772-115-311_B10 |  |
 |
 |
| 1772-115-311_B11 |  |
 |
 |
| 1772-115-311_B12 |  |
 |
 |
| 1772-115-311_C01 |  |
 |
 |
| 1772-115-311_C02 |  |
 |
 |
| 1772-115-311_C03 |  |
 |
 |
| 1772-115-311_C04 |  |
 |
 |
| 1772-115-311_C05 |  |
 |
 |
| 1772-115-311_C06 |  |
 |
 |
| 1772-115-311_C07 |  |
 |
 |
| 1772-115-311_C08 |  |
 |
 |
| 1772-115-311_C09 |  |
 |
 |
| 1772-115-311_C10 |  |
 |
 |
| 1772-115-311_C11 |  |
 |
 |
| 1772-115-311_C12 |  |
 |
 |
| 1772-115-311_D01 |  |
 |
 |
| 1772-115-311_D02 |  |
 |
 |
| 1772-115-311_D03 |  |
 |
 |
| 1772-115-311_D04 |  |
 |
 |
| 1772-115-311_D05 |  |
 |
 |
| 1772-115-311_D06 |  |
 |
 |
| 1772-115-311_D07 |  |
 |
 |
| 1772-115-311_D08 |  |
 |
 |
| 1772-115-311_D09 |  |
 |
 |
| 1772-115-311_D10 |  |
 |
 |
| 1772-115-311_D11 |  |
 |
 |
| 1772-115-311_D12 |  |
 |
 |
| 1772-115-311_E01 |  |
 |
 |
| 1772-115-311_E02 |  |
 |
 |
| 1772-115-311_E03 |  |
 |
 |
| 1772-115-311_E04 |  |
 |
 |
| 1772-115-311_E05 |  |
 |
 |
| 1772-115-311_E06 |  |
 |
 |
| 1772-115-311_E07 |  |
 |
 |
| 1772-115-311_E08 |  |
 |
 |
| 1772-115-311_E09 |  |
 |
 |
| 1772-115-311_E10 |  |
 |
 |
| 1772-115-311_E11 |  |
 |
 |
| 1772-115-311_E12 |  |
 |
 |
| 1772-115-311_F01 |  |
 |
 |
| 1772-115-311_F02 |  |
 |
 |
| 1772-115-311_F03 |  |
 |
 |
| 1772-115-311_F04 |  |
 |
 |
| 1772-115-311_F05 |  |
 |
 |
| 1772-115-311_F06 |  |
 |
 |
| 1772-115-311_F07 |  |
 |
 |
| 1772-115-311_F08 |  |
 |
 |
| 1772-115-311_F09 |  |
 |
 |
| 1772-115-311_F10 |  |
 |
 |
| 1772-115-311_F11 |  |
 |
 |
| 1772-115-311_F12 |  |
 |
 |
| 1772-115-311_G01 |  |
 |
 |
| 1772-115-311_G02 |  |
 |
 |
| 1772-115-311_G03 |  |
 |
 |
| 1772-115-311_G04 |  |
 |
 |
| 1772-115-311_G05 |  |
 |
 |
| 1772-115-311_G06 |  |
 |
 |
| 1772-115-311_G07 |  |
 |
 |
| 1772-115-311_G08 |  |
 |
 |
| 1772-115-311_G09 |  |
 |
 |
| 1772-115-311_G10 |  |
 |
 |
| 1772-115-311_G11 |  |
 |
 |
| 1772-115-311_G12 |  |
 |
 |
| 1772-115-311_H01 |  |
 |
 |
| 1772-115-311_H02 |  |
 |
 |
| 1772-115-311_H03 |  |
 |
 |
| 1772-115-311_H04 |  |
 |
 |
| 1772-115-311_H05 |  |
 |
 |
| 1772-115-311_H06 |  |
 |
 |
| 1772-115-311_H07 |  |
 |
 |
| 1772-115-311_H08 |  |
 |
 |
| 1772-115-311_H09 |  |
 |
 |
| 1772-115-311_H10 |  |
 |
 |
| 1772-115-311_H11 |  |
 |
 |
| 1772-115-311_H12 |  |
 |
 |
| 1772-115-314_A01 |  |
 |
 |
| 1772-115-314_A02 |  |
 |
 |
| 1772-115-314_A03 |  |
 |
 |
| 1772-115-314_A04 |  |
 |
 |
| 1772-115-314_A05 |  |
 |
 |
| 1772-115-314_A06 |  |
 |
 |
| 1772-115-314_A07 |  |
 |
 |
| 1772-115-314_A08 |  |
 |
 |
| 1772-115-314_A09 |  |
 |
 |
| 1772-115-314_A10 |  |
 |
 |
| 1772-115-314_A11 |  |
 |
 |
| 1772-115-314_A12 |  |
 |
 |
| 1772-115-314_B01 |  |
 |
 |
| 1772-115-314_B02 |  |
 |
 |
| 1772-115-314_B03 |  |
 |
 |
| 1772-115-314_B04 |  |
 |
 |
| 1772-115-314_B05 |  |
 |
 |
| 1772-115-314_B06 |  |
 |
 |
| 1772-115-314_B07 |  |
 |
 |
| 1772-115-314_B08 |  |
 |
 |
| 1772-115-314_B09 |  |
 |
 |
| 1772-115-314_B10 |  |
 |
 |
| 1772-115-314_B11 |  |
 |
 |
| 1772-115-314_B12 |  |
 |
 |
| 1772-115-314_C01 |  |
 |
 |
| 1772-115-314_C02 |  |
 |
 |
| 1772-115-314_C03 |  |
 |
 |
| 1772-115-314_C04 |  |
 |
 |
| 1772-115-314_C05 |  |
 |
 |
| 1772-115-314_C06 |  |
 |
 |
| 1772-115-314_C07 |  |
 |
 |
| 1772-115-314_C08 |  |
 |
 |
| 1772-115-314_C09 |  |
 |
 |
| 1772-115-314_C10 |  |
 |
 |
| 1772-115-314_C11 |  |
 |
 |
| 1772-115-314_C12 |  |
 |
 |
| 1772-115-314_D01 |  |
 |
 |
| 1772-115-314_D02 |  |
 |
 |
| 1772-115-314_D03 |  |
 |
 |
| 1772-115-314_D04 |  |
 |
 |
| 1772-115-314_D05 |  |
 |
 |
| 1772-115-314_D06 |  |
 |
 |
| 1772-115-314_D07 |  |
 |
 |
| 1772-115-314_D08 |  |
 |
 |
| 1772-115-314_D09 |  |
 |
 |
| 1772-115-314_D10 |  |
 |
 |
| 1772-115-314_D11 |  |
 |
 |
| 1772-115-314_D12 |  |
 |
 |
| 1772-115-314_E01 |  |
 |
 |
| 1772-115-314_E02 |  |
 |
 |
| 1772-115-314_E03 |  |
 |
 |
| 1772-115-314_E04 |  |
 |
 |
| 1772-115-314_E05 |  |
 |
 |
| 1772-115-314_E06 |  |
 |
 |
| 1772-115-314_E07 |  |
 |
 |
| 1772-115-314_E08 |  |
 |
 |
| 1772-115-314_E09 |  |
 |
 |
| 1772-115-314_E10 |  |
 |
 |
| 1772-115-314_E11 |  |
 |
 |
| 1772-115-314_E12 |  |
 |
 |
| 1772-115-314_F01 |  |
 |
 |
| 1772-115-314_F02 |  |
 |
 |
| 1772-115-314_F03 |  |
 |
 |
| 1772-115-314_F04 |  |
 |
 |
| 1772-115-314_F05 |  |
 |
 |
| 1772-115-314_F06 |  |
 |
 |
| 1772-115-314_F07 |  |
 |
 |
| 1772-115-314_F08 |  |
 |
 |
| 1772-115-314_F09 |  |
 |
 |
| 1772-115-314_F10 |  |
 |
 |
| 1772-115-314_F11 |  |
 |
 |
| 1772-115-314_F12 |  |
 |
 |
| 1772-115-314_G01 |  |
 |
 |
| 1772-115-314_G02 |  |
 |
 |
| 1772-115-314_G03 |  |
 |
 |
| 1772-115-314_G04 |  |
 |
 |
| 1772-115-314_G05 |  |
 |
 |
| 1772-115-314_G06 |  |
 |
 |
| 1772-115-314_G07 |  |
 |
 |
| 1772-115-314_G08 |  |
 |
 |
| 1772-115-314_G09 |  |
 |
 |
| 1772-115-314_G10 |  |
 |
 |
| 1772-115-314_G11 |  |
 |
 |
| 1772-115-314_G12 |  |
 |
 |
| 1772-115-314_H01 |  |
 |
 |
| 1772-115-314_H02 |  |
 |
 |
| 1772-115-314_H03 |  |
 |
 |
| 1772-115-314_H04 |  |
 |
 |
| 1772-115-314_H05 |  |
 |
 |
| 1772-115-314_H06 |  |
 |
 |
| 1772-115-314_H07 |  |
 |
 |
| 1772-115-314_H08 |  |
 |
 |
| 1772-115-314_H09 |  |
 |
 |
| 1772-115-314_H10 |  |
 |
 |
| 1772-115-314_H11 |  |
 |
 |
| 1772-115-314_H12 |  |
 |
 |
InCell Analyzer 6000 Z Stack Movies
| Cell ID | Brightfield Z-Stack Video | Green Fluorescence Z-Stack Video | Red Fluorescence Z-Stack Video |
|---|
Technical Metadata
| Metadata ID | Cell Line | No. of Samples | No. of Images | Sequencer | Material | Capture |
|---|---|---|---|---|---|---|
| 0003 | A549 | 1344 | 4032 | Illumina HiSeq 2500 | Adenocarcinomic human alveolar basal epithelial cells | C1 Single-cell Auto Prep Integrated Fluidic Circuits (IFC) |